The CGAT AlignmentViewer program
Starting AlignmentViewer
On Windows or Mac OSX, double clicking on the "AlignmentViewer" icon invokes the program. On Unix systems, type the following command on the command line:
java -jar AlignmentViewer.jar |
However, when invoking with the above command, you will be likely to encounter a memory allocation error especially when you want to see an alignment of long sequences generated by dynamical calculation. To avoid this, we recommend you to increase the size of the memory allocation pool for the Java virtual machine with the -Xmx option.
java -Xmx512m -jar AlignmentViewer.jar |
Load data
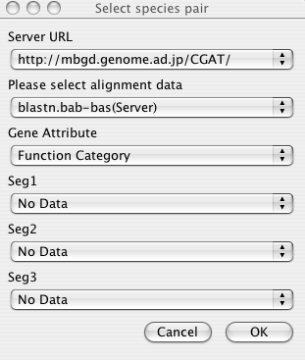
After running the program, first you must load data by choosing File ⇒ Load from the menu.
- Server URL
Choose the server from which data are downloaded. By default, only the default CGAT server (running on the MBGD server) is registered. You can add more servers to this list in the Locations section of the Properties panel (File ⇒ Properties), if you or your colleagues have been setup an original new server. See the chapter " The CGAT DataServer package " for server setup and the " Configuring AlignmentViewer "section for properties setup.
- Alignment Data
Choose the alignment data to be downloaded specified by the combination of organism names and a program name. Note that choosing the server URL above reset the list of available alignment data. If you have setup the data on the local file system that the AlignmentViewer program can directly accessed, and have correctly set the CGAT_HOME variable in the Properties panel, you can also choose the data available on the local machine. These data are listed without the "(Server)" indication.
- Gene Attribute
Choose the gene attribute data for coloring genes.
- Feature Segment Data
Choose the feature segment data to be shown on the additional annotation tracks (see the section "Alignment display panel "). By default, maximally three types of segment data can be loaded at once, but the maximum number can be increased in the Properties panel.