Install
AlignmentViewer
Basically, you need not particular operation after having downloaded the program file (AlignmentViewer.jar) — double click the icon (OSX or Windows) or type the java command (Unix) to invoke the program. See the chapter " The CGAT AlignmentViewer program "
However, in the default setting of the Java virtual machine (JVM), you will be likely to encounter a memory allocation error especially when aligning long sequences dynamically. To avoid this problem, you need to increase the size of the memory allocation pool for the JVM.
Windows: Download the AlignmentViewer.jar file and do the following (see Figure 1).
Create a shortcut of the AlignmentViewer.jar file (Right-click on the jar file ⇒ Create Shortcut).
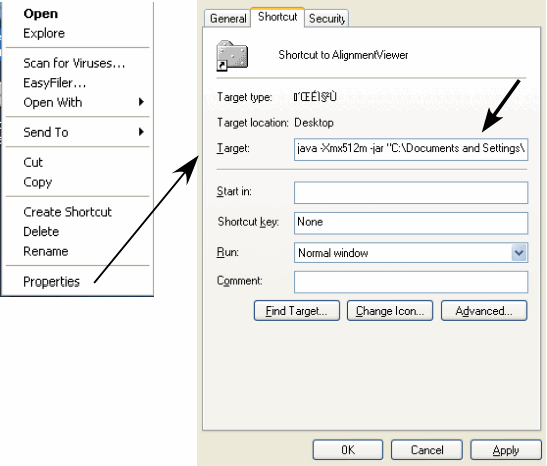
Open the properties window of the shortcut (Right-click on the shortcut ⇒ Properties).
Open the Shortcut tab and enter the java command in the Target: box as follows: [ java -Xmx512m -jar "C:\Documents and Settings\..." ], where "C:\Documents and Settings\..." is the original file location. This might be different in you situation, but do not modify the original name in anyway. Then, press "OK".
Mac OSX: Download and use the OSX application version of AlignmentViewer (AlignmentViewer.app), instead of the AlignmentViewer.jar file.
Linux/Unix: Download the AlignmentViewer.jar file and type java -Xmx512m -jar AlignmentViewer.jar to invoke the program.
DataServer
Copy the downloaded archive CGAT.tgz into an appropriate directory and extract files by tar command, which generates a directory named CGAT. Enter this directory and invoke a configuration script SetUp.pl.
tar cvfz CGAT.tgz cd CGAT SetUp.pl |
# in csh or tcsh source etc/cgat.csh # in sh or bash . etc/cgat.sh |
The set up script tries to detect alignment programs available on the server machine and installs detected programs in the $CGAT_HOME/build/align directory. You might want to move some commands from the $CGAT_HOME/build/align.in directory or to remove some unnecessary commands manually.
Similarly, you might want to install manually some optional commands for collecting feature segments from the $CGAT_HOME/build/segment.opt directory. Note that in this case you must always set up by yourself (if you wanted), since currently the set up script does not try to check the availability of these commands automatically. Currently, the following commands are available:
Glimmer: Microbial gene-finding system available at http://www.cbcb.umd.edu/software/glimmer/. Currently, ver.2 is assumed by default, but you can use ver.3 if you manually edit the commands/Glimmer.pl file. Before that, of course, you must correctly set up the glimmer program itself.
ISfind: Searching an insertion sequence (IS) database by the BLAST program. The IS database should be a FASTA formatted sequence file with each comment line containing IS family name as ">${ISname} Fam:${familyname}", and the file should be installed as $CGAT_HOME/database/isdb . See the IS Finder database for information about IS families. One can use the GIB-IS database available at DDBJ as a downloadable IS database. Download the table file (IS-table-text.txt) and run the following command:
$CGAT_HOME/build/bin/conv_GIB_IS.pl -nr IS-table-text.txt > $CGAT_HOME/database/isdb