MBGD Top > Quick Tour > Search
Searching against an orthologous gene table
- On the top page, there is a table containing a list of all available
organisms. Once you have changed a set of target organisms,
only selected organisms are highlighted in green here (you might need to press
"Reload" button on your browser
for the correct display). You can change the target organisms
by pressing "Taxonomy Browser" button.
Notice that when you change the target organsisms here, a clustering
procedure may be invoked when required.
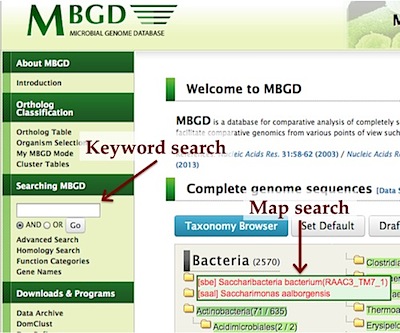
From the top page, you can search the database by several methods:- Keyword search on the top page just searches for entered terms against gene records.
- Advanced search enables you to specify keywords for each field individually.
- Homology search searches your query sequences against protein sequences in MBGD.
- Map search enables you to navigate chromosome map of individual genome to retrieve a particular gene.
Now, we've selected some target organisms at the tour1.
- Enter keywords and press return.
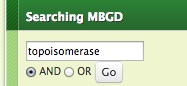
MySQL fulltext index is used for this search. You can switch how to combine the multiple words specified (AND/OR) using the radio button below.
- The result is returned. Click the [Show Cluster Table] button in the bottom of the result page.
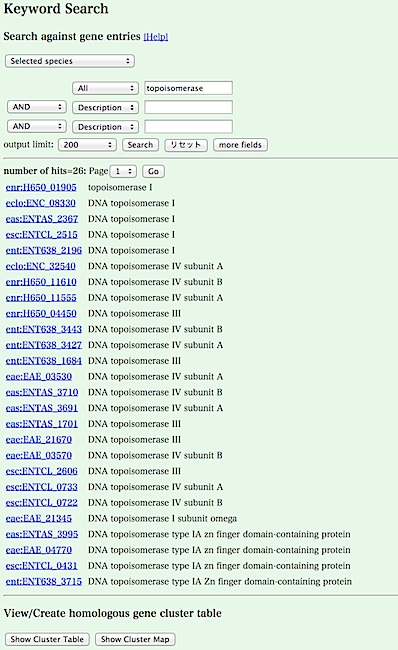
- You can see a cluster table containing the retrieved genes. Here, the table shows a pattern of presence/absence of ortholgs in each genome for each orthlogous group (i.e. phylogenetic pattern).
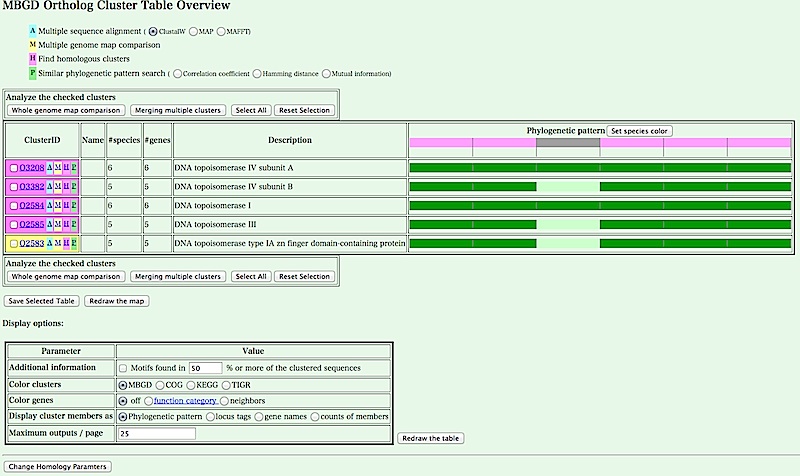
You can change the display of the table using Display Options below. For example, if you select 'locus tags' in 'Display cluster members as', and click [Redraw the table] button, you can see a table with a locus tag for each gene, where genes hit by the keyword search are highlighted in red.
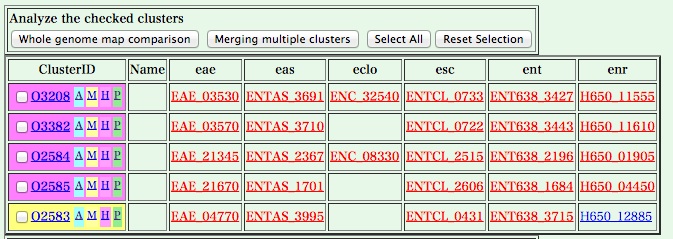
Click a locus tag to retireve a gene record.
- Gene information page is displayed.
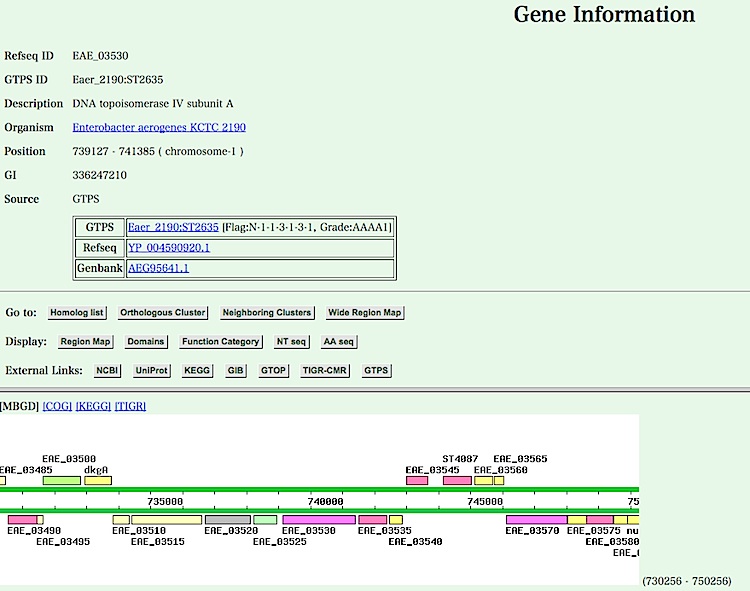
You can see several information about this gene by clicking buttons listed in the upper frame. Now, click the "Neighboring Clusters" button.
- In this table, orthologous clusters are arranged in the order of the
reference genome (in this example, Enterobacter aerogenes (eae) genome), and
genes that are adjacent to each other are indicated in the same colors.
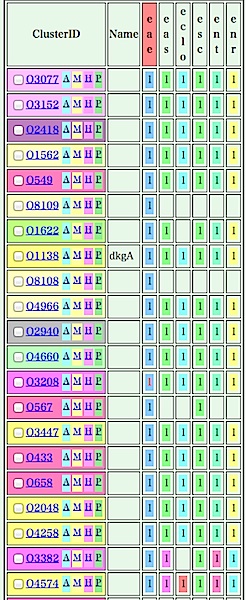
Let's move to the next tour: 3. Phylogenetic pattern analysis