MBGD Top > Quick Tour > Phylogenetic pattern analysis
Phylogenetic pattern analysis
-
Phylogenetic pattern (or phylogenetic profile) is a binary vector defined for each orthologous group that represents presence or absence of that orthologs in each genome.
It is known that orthologous groups having similar phylogenetic patterns are often functionally related to each other, and finding similar phylogenetic patterns is one of the effective approaches for functional prediciton through comparative genomics.
MBGD provides users a function to search for orthologous groups with similar phylogenetic patterns to a specified orthologous group. MBGD also allows users to specify a phylogenetic pattern directly to search for genes with similar phylogenetic patterns.
In the following example, we will speciy a phylogenetic pattern related to some phynotypic property and search for orthologouos groups related to this phylogenetic pattern.
First of all, click "Set Default" in the top page to clear the set of target organisms.
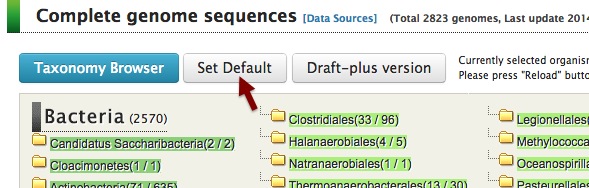
Then click "Ortholog Table" in the left menu of the top page.
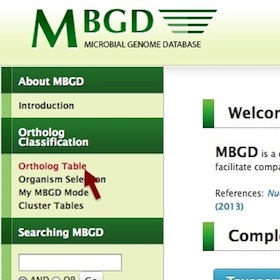
-
Scroll to bottom, the clustering summary page is displayed.
Find the "serch conditions" box below the cluster size histogram.
Press [Change pattern] button in the occurrence pattern section to specify phylogenetic pattern to search for orthologous groups.
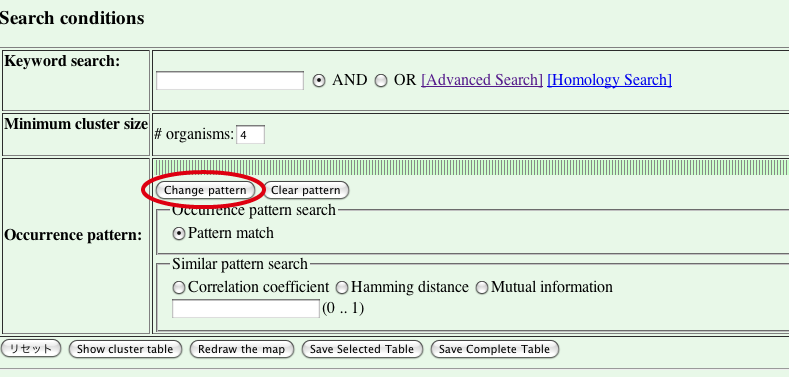
- Organism selection page is displayed.
In this interface, you can specify "present organsisms" and "absent organsisms", which are sets of organisms in which the orthologs in question are present and absent, repsectively. Basically, you can choose a set of organisms in the central box and press [Add to presence] or [Add to absense] to add the organisms to either set. You can also use the phenotypic information from the GOLD database in the right panel of this interface.
For this purpose, first you should choose a property type listed on the top of the right panel.
Here we choose "motility" as a property and specify "Motile" for its value. Then, the set of organisms shown in the central panel is redisplayed, showing only motile organisms.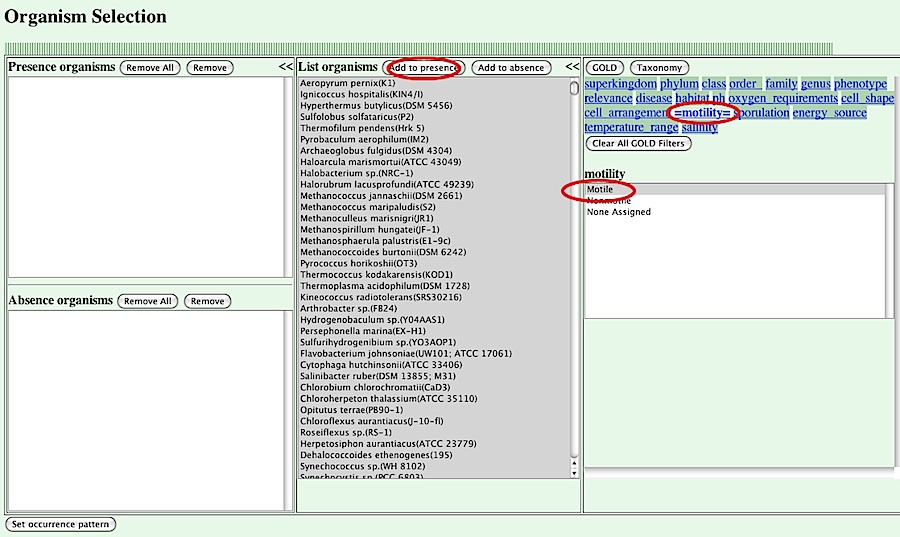
Press [Add to presence] button in the central panel to add the set of organisms into the list of present organsims in the left panel. Next, choose "Nonmotile" in the right panel to choose nonmotile organisms and press [Add to absence] in the central panel to specify absent organisms.
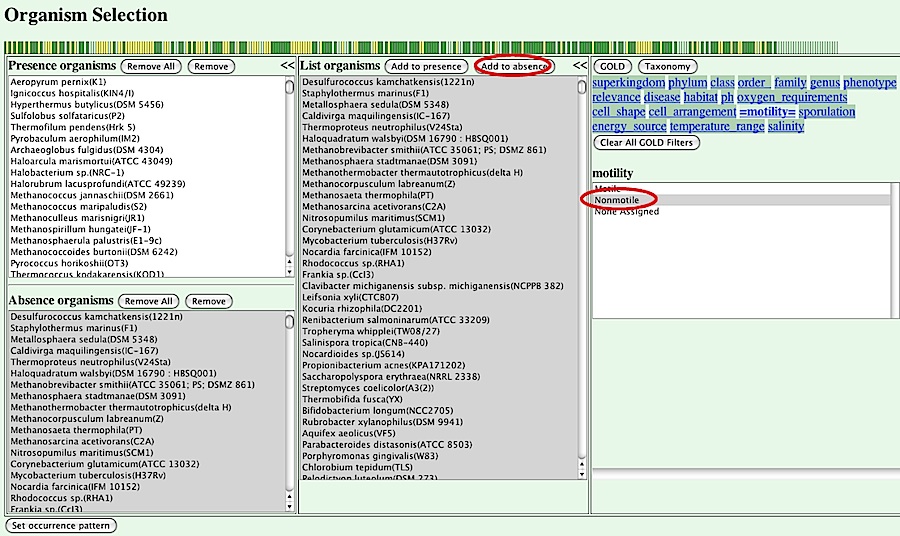
The resulting phylogenetic pattern is displayed graphically at the top of this page, where present organisms are shown in green and absent organisms are shown in yellow. In this case, the pattern represents genes present in the motile organsisms and absent in the nonmotile organisms.
Then, press [Set occurrence pattern] button to finish the specification.
- The search condition panel is now redisplayed.
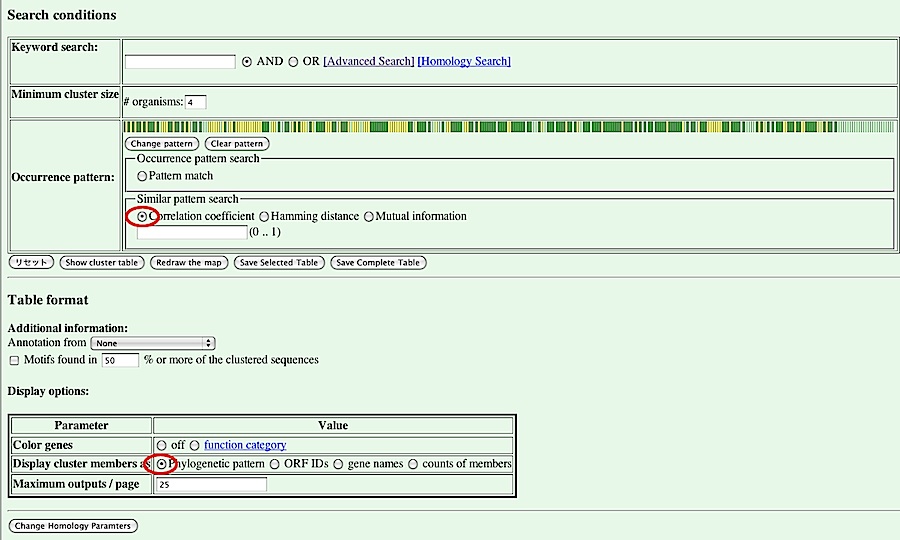
To search for orthologous groups having similar phylogenetic pattern to the specified one, you should choose a similarity measure from the list including Correlation coefficient, Hamming distance, and Mutial information. Here we choose "Correlation coefficient". You should also check that "Display cluster members as" in the Display option box is "Phylogenetic pattern". Then, press [Show cluster table] button.
- The cluster table is displayed, showing orthologous groups related to the above specified "motility specific phylogenetic pattern".
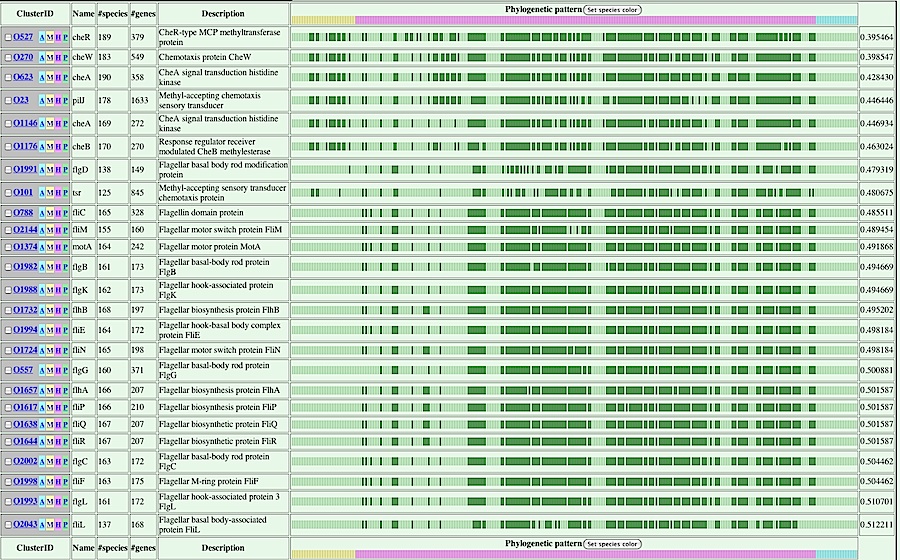
Here, the table is ordered according to the dissimilarity value calculated based on the correlation coefficient between the specified phylogenetc pattern and that of each ortholog group. Phylogenetic patterns are graphically displayed with green bars indicating "present". The dissimilarity values, which are defined here as d=(1-r)/2, where r is correlation coefficient, are shown in the right most column.
By clicking the [P] button in the first collumn of an appropriate ortholog group, you can re-order the cluster table according to the similarity of phylogenetic patterns between the specified cluster and each cluster.
Let's move to the next tour: 4. Create ortholog table.