MBGD Top > Quick Tour > MyMBGD (+draft genome version)
My MBGD: Add draft genomes to your ortholog analysis in MBGD
-
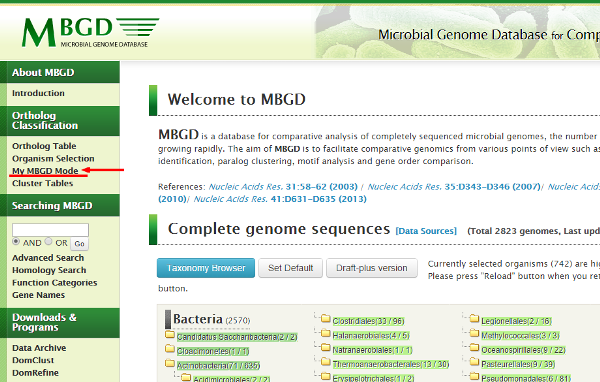
To analyze genome including draft, firstly you must enter the "My MBGD Mode" from
the top page menu.
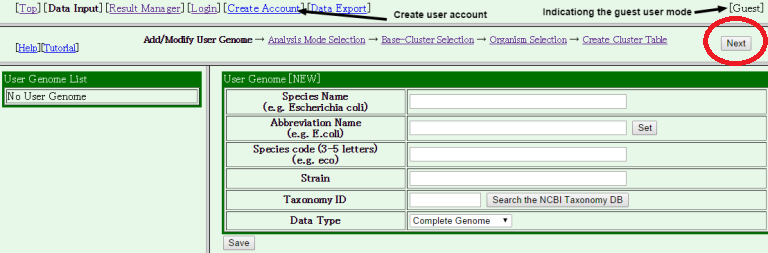
- The data input form is displayed. If you do not register your data, you can skip this process by clicking "Next"
- Next, choose an analysis mode.
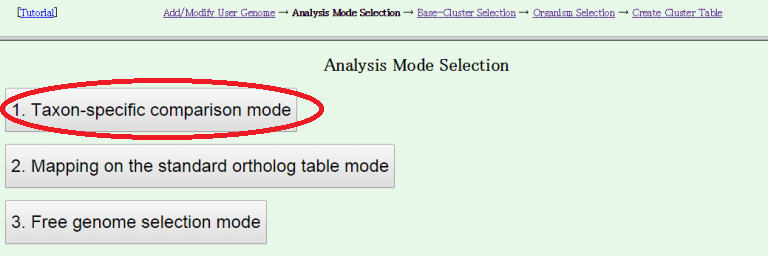
Users can choose from the following three modes of analysis:
- Taxon specific comparison mode
First choose a taxon to be analyzed and then specify a set of genomes to be compared within that taxon. In this mode, users can choose either from scratch or adding to a taxon-specific ortholog table. - Mapping on the standard ortholog table mode
Selected genomes will be added to the standard ortholog table using MergeTree. - Free genome selection mode, which is equivalent to the previous MyMBGD mode
Users can freely choose a set of genomes to compare including user genomes, complete genomes and draft genomes and conduct ortholog analysis using DomClust.
Here, we choose "Taxon-specific comparisom mode"
- Taxon specific comparison mode
- Next, choose a taxon to be analyzed.
Here, we choose "Staphylococcus aureus (species)" as a base cluster.

- Next, choose genome sequences to be compared.
The maximum number of genomes to be selected is limited according
to the size of your data.
In this case, you can choose maximally 50 genomes.
By default, no draft genome is included in this list. To view draft genomes, please click "Show Drafts (Reload)".
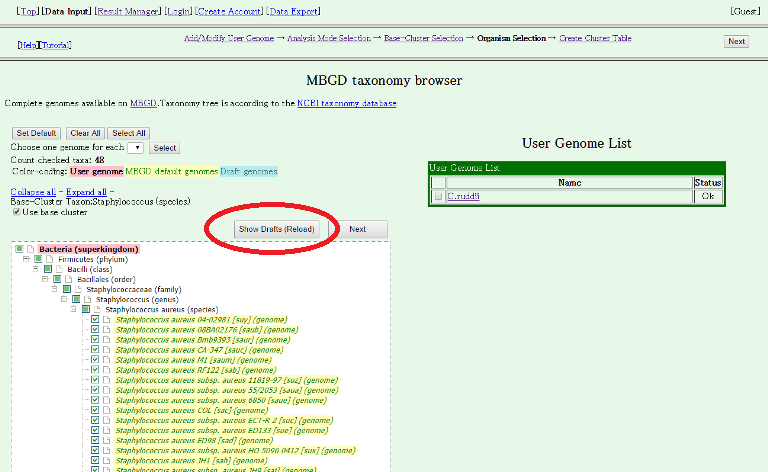
Then, draft genomes are displayed.
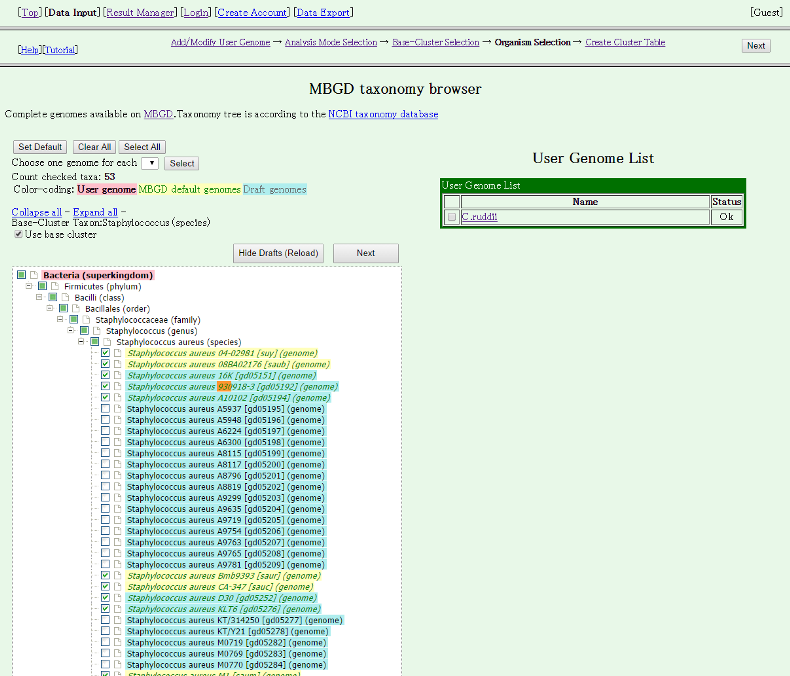
Here, we choose five draft genomes (strains 16K, 930918-3, A10102, D30 and KLT6) in Staphylococcus aureus. Note that "Use base cluster" checkbox is checked, which indicates that the genes in the selected draft genomes will be added to the existing ortholog table of Staphylococcus aureus incrementally. Thus, total 53 genomes including 48 complete genomes of the base cluster and 5 draft genomes to be added are selected to compare.
- The Homologous Gene Table Ceration page is displayed. Press "Create/View Cluster Table" to start building.
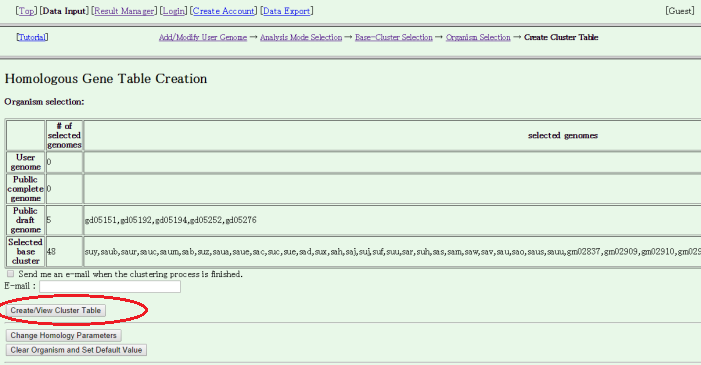
The building process includes database construction, all-all homology search, motif search and orthlog clustering. In this case with relatively small data, probably you should wait several minutes until the process is completed, but it may take more time if the server is busy.
Please be patient!
- You are done! The clustering result is displayed.
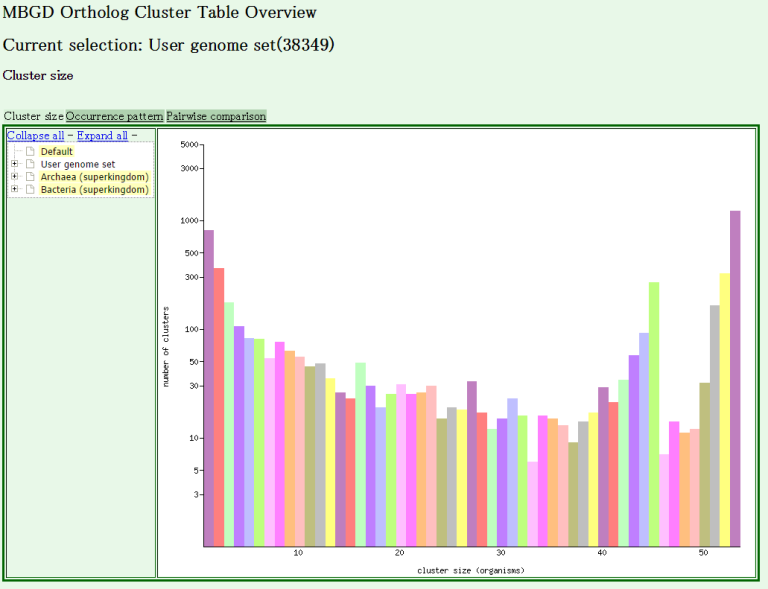
- You can use every function of MBGD as in the usual mode.
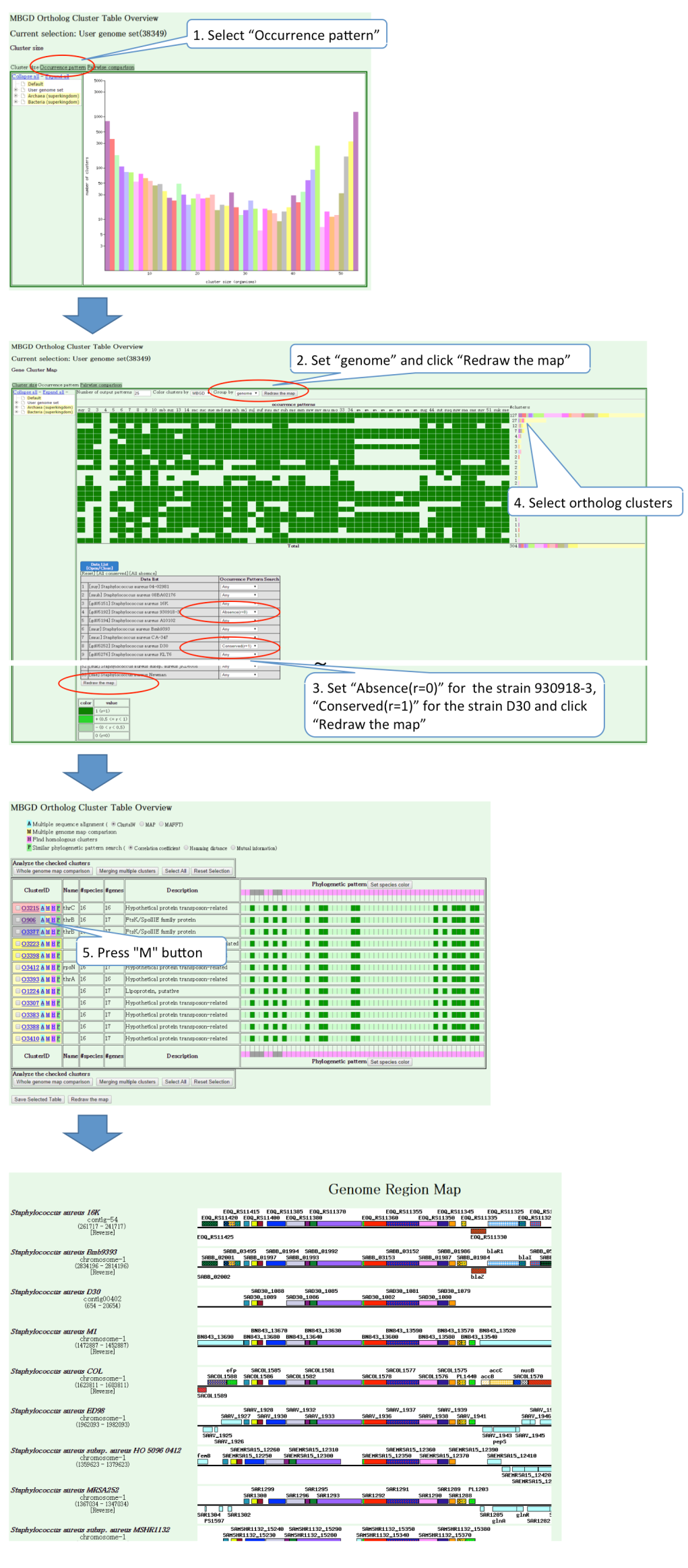
The following is a sample session to analyze the clustering result. In this example, the strains D30 and 930918-3 are models for nasal carrier and non-carrier strains, respectively (Sivaraman, K. et al, BMC Genomics, 9, 433). To elucidate the difference between these strains, the occurrence pattern analysis is carried out (Steps 1,2). After selecting the occurrence patterns satisfying the condition "present in D30 and absent in 930918-3" (Step 3), choose a characteristic occurrence pattern where genes are present only in some specific strains (Step 4) to display the ortholog groups showing this pattern. The resulting table suggests that they are likely to be part of a transposon. By clicking the "M" button in one of the leftmost cells, you can see a comparative chromsome map around this ortholog group and confirm that these genes are indeed clustered on each chromosome (Step 5).
Let's move to the next tour: 6. My MBGD: Add your own genome data to MBGD.